用ggplot2在分组条形图中添加不同列的错误条
用ggplot2在分组条形图中添加不同列的错误条
提问于 2020-12-13 19:08:44
我想把错误条添加到我的酒吧里。标准偏差的数据在两个不同的列中。
这是我的酒馆:
# load packages
> library(data.table)
> library(ggplot2)
> library(tidyr)
>
> results <- fread("Results.csv", header=TRUE, sep=";")
>
> str(results)
Classes ‘data.table’ and 'data.frame': 7 obs. of 5 variables:
$ Organism : chr "AC1432" "D3425" "BF3523" "XR2405" ...
$ Molecule1 : num 39.5 418.4 189.2 49.3 4610.9 ...
$ Molecule1sd: num 19.6 70.9 102.8 21.2 275.9 ...
$ Molecule2 : num 276 6511 235 500 11205 ...
$ Molecule2sd: num 21 291.1 109.7 67.1 94.5 ...
- attr(*, ".internal.selfref")=<externalptr>
>
> df <- data.frame(results)
>
> str(df)
'data.frame': 7 obs. of 5 variables:
$ Organism : chr "AC1432" "D3425" "BF3523" "XR2405" ...
$ Molecule1 : num 39.5 418.4 189.2 49.3 4610.9 ...
$ Molecule1sd: num 19.6 70.9 102.8 21.2 275.9 ...
$ Molecule2 : num 276 6511 235 500 11205 ...
$ Molecule2sd: num 21 291.1 109.7 67.1 94.5 ...
>
>
>
> # Manually set factor levels of 'Organism' column to plot in a logical order.
> df$Organism = factor(df$Organism,
+ levels=c("without organism", "AC1432", "BF3523", "XR2405", "D3425", "XR2463", "ATF259"))
>
> df.g <- gather(df, Molecule1, Molecule2, -Organism, -Molecule1sd, -Molecule2sd)
> df.sd <- gather(df, Molecule1sd, Molecule2sd, -Molecule1, -Molecule2, -Organism)
> ggplot(df.g, aes(Molecule1, Molecule2)) +
+ geom_bar(aes(fill = Organism), stat = "identity", position = "dodge")使用的数据:
> dput(df)
structure(list(Organism = structure(c(2L, 5L, 3L, 4L, 6L, 7L,
1L), .Label = c("without organism", "AC1432", "BF3523", "XR2405",
"D3425", "XR2463", "ATF259"), class = "factor"), Molecule1 = c(39.45920899,
418.4234805, 189.162295, 49.314698, 4610.921188, 751.7070352,
35), Molecule1sd = c(19.55450482, 70.91013667, 102.7566193, 21.20841393,
275.8934527, 71.62450643, NA), Molecule2 = c(275.9147606, 6510.974605,
235.247381, 499.8928585, 11205.33907, 9507.869294, 250), Molecule2sd = c(21.04668977,
291.1223384, 109.652064, 67.1000078, 94.54544271, 707.1950335,
NA)), row.names = c(NA, -7L), class = "data.frame")这是我对错误栏的尝试
ggplot(df.g, aes(Molecule1, Molecule2)) +
geom_bar(aes(fill = Organism), stat = "identity", position = "dodge") +geom_errorbar(df.sd, aes_Molecule1(ymin=Molecule1-Molecule1sd, ymax=Molecule1+Molecule1sd),aes_Molecule2(ymin=Molecule2-Molecule2sd, ymax=Molecule2+Molecule2sd), width=.2 )但我的主意行不通。如何从两个不同的列中添加错误条?
回答 1
Stack Overflow用户
回答已采纳
发布于 2020-12-13 20:36:44
如果您使用有机体、分子、平均值和sd的列重新塑造数据集,可能会更容易。下面是一种tidyverse方法:
包和数据集
library(tidyverse)
df <- data.frame(Organism = c("AC1432", "D3425", "BF3523", "XR2405",
"XR2463", "ATF259", "without organism"),
Molecule1 = c(39.5, 418.4, 189.2, 49.3,
4610.9, 800, 10),
Molecule1sd = c(19.6, 70.9, 102.8, 21.2,
275.9, 100, 1),
Molecule2 = c(276, 6511, 235, 500,
11205, 9500, 250),
Molecule2sd = c( 21, 291.1, 109.7, 67.1,
94.5, 50, 2))
# I estimated the not shown values in your str(result)重塑
df2 <- df %>%
# add meaningful ending to columnnames containing mean (m)
select(Molecule1m = Molecule1,
Molecule2m = Molecule2,
everything()) %>%
# gather whole dataset into Molecule, mean, sd
pivot_longer(cols = -Organism,
names_to = c("Molecule", ".value"),
names_pattern = "(Molecule[12])(.)") %>%
# factor reorder levels
mutate(Organism = factor(Organism,
levels=c("without organism", "AC1432",
"BF3523", "XR2405",
"D3425", "XR2463", "ATF259")))图
ggplot(df2, aes(x = Molecule,
y = m,
fill = Organism)) +
geom_col(position = "dodge") +
geom_errorbar(aes(ymin = m - s, ymax = m + s),
position = "dodge")
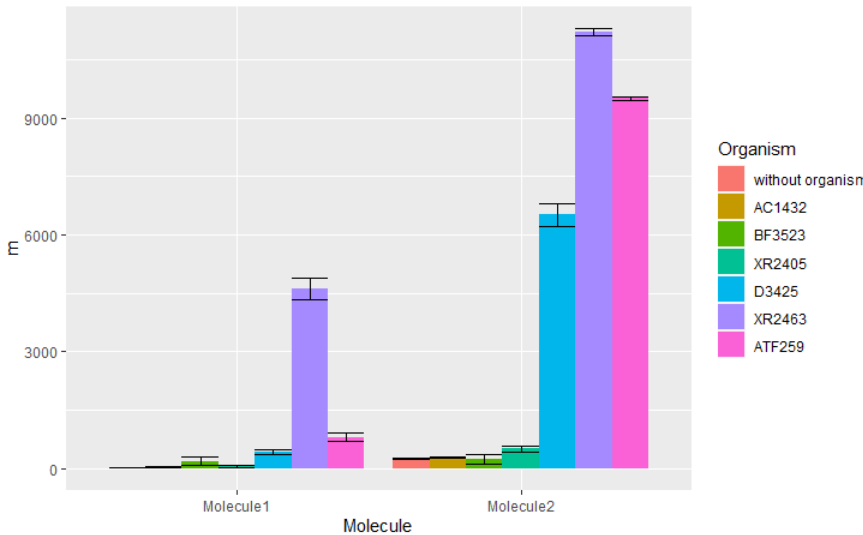
页面原文内容由Stack Overflow提供。腾讯云小微IT领域专用引擎提供翻译支持
原文链接:
https://stackoverflow.com/questions/65279545
复制相关文章
相似问题

