【一步到位】统一空转visium以及scRNA单细胞seurat转h5ad
【一步到位】统一空转visium以及scRNA单细胞seurat转h5ad

KS科研分享与服务-TS的美梦
发布于 2026-04-02 13:04:01
发布于 2026-04-02 13:04:01
之前介绍过空转h5ad转seurat的操作(空转python版【2】:visium空转h5ad数据转化为seurat测试)。类似于scRNA一样,小伙伴对于空转seurat转h5ad有很大的需求。但是网上没有直接的方式或者说简洁的办法能够轻松实现空转数据的转化。这里以visium空转为例,介绍两种方式,第一种是比较迂回的路线,也算是后续函数的思路。第二种方式整合了一个函数,能够一步到位实现空转visium seurat到h5ad的转化,且相当于统一了scRNA以及空转的转化,次函数也能实现普通房scRNA数据seurat转化为h5ad。对于空转,函数只适用于单个sample的转化!
首先看看示例数据:
#R中演示
#----------------------------------------------------------------------------------
#我们这里演示的visium数据包含多个样本,提取单个的进行演示
#多个的数据其实也是需要先分开在进行转化的
library(Seurat)
CTR3 <- subset(merged,orig.ident=='CTR3')
#查看整合的降维UMAP
color_pal<-c("#11A579","#F2B701","#66C5CC","#80BA5A","#F6CF71","#7F3C8D","#CF1C90","#3969AC","#f97b72","#E73F74","#4b4b8f","#ACA4E2","#31C53F","#B95FBB","#D4D915","#28CECA")
DimPlot(CTR3, reduction = "umap", label = F, pt.size = 0.5,cols = color_pal)
#可视化slice
SpatialDimPlot(CTR3, stroke=0.1)& #如果spots不需要边框,设置stroke=NA
scale_fill_manual(values = color_pal)
#----------------------------------------------------------------------------------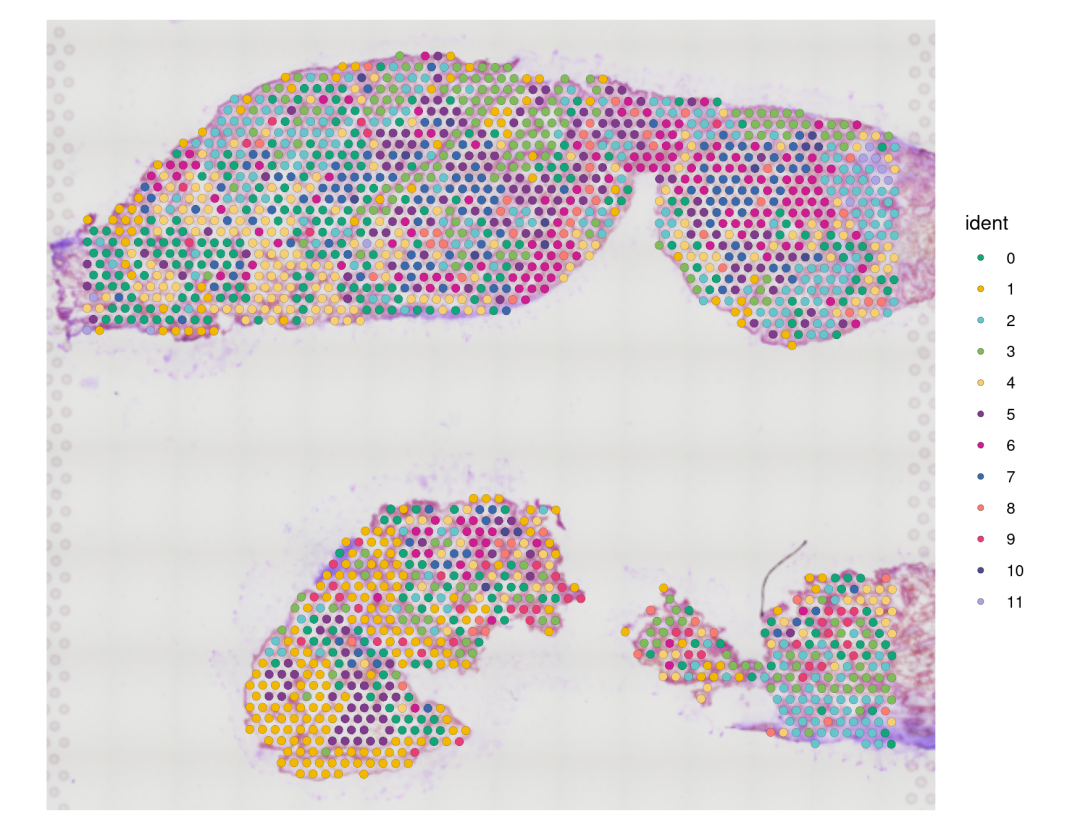
image.png
第一种方式:
迂回路线,目前(2025年底)我是没有看到较好的方式能够轻松实现空转seurat转化为h5ad,很多时候空转不太需要转化,直接使用python读入即可,需要转化是因为我们已经在R中进行了数据处理,并且需要保持这些内容在python中进行分析。
关于seurat转h5ad有很多种方式(比如sceasy,anndata,或者我们合集中介绍的其他方式,还有一些我没有接触过的包),但是不能保留图片信息,所以最简单采取的路线就是先转化一个没有image信息的对象,然后添加图片信息进行就可以了!参考python读入空转数据:空转python版【1】:visium分析(scanpy+squidpy)---矩阵只有10X标准3个文件怎么办?
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import anndata as ad
import scanpy as sc
import squidpy as sq#这里我是直接使用转化好的演示,转化的数据只有矩阵信息,没有image
CTR3 = sc.read_h5ad('./CTR3.h5ad')
CTR3
# AnnData object with n_obs × n_vars = 1801 × 18274
# obs: 'orig.ident', 'nCount_Spatial', 'nFeature_Spatial', 'group', 'percent.mt', 'nCount_SCT', 'nFeature_SCT', 'nCount_RNA', 'nFeature_RNA', 'integrated_snn_res.0.6', 'seurat_clusters'
# obsm: 'pca', 'umap'sc.pl.umap(CTR3,color='seurat_clusters')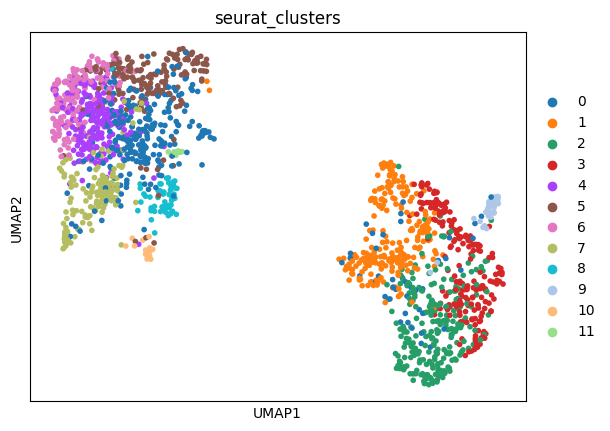
No description has been provided for this image
#因为之前的数据分析是整合的,所以原有的barcode加了后缀,为了与原始的一致,需要处理一下,把多余添加的后缀去除
#这里的处理是直接使用replace将“_1“替换掉就可以了
CTR3.obs_names = CTR3.obs_names.str.replace(r'_1$', '', regex=True)#读入tissue_positions文件并过滤在组织上的spots
pos = pd.read_csv('./visium/CTR3_processed_data/spatial/tissue_positions.csv')
pos = pos.set_index("barcode")
pos = pos.loc[pos.index.isin(CTR3.obs_names)]#添加坐标
CTR3.obs = CTR3.obs.merge(pos, left_index=True, right_index=True, how="left")
pxr, pxc = "pxl_row_in_fullres", "pxl_col_in_fullres"
CTR3.obsm["spatial"] = CTR3.obs[[pxc, pxr]].to_numpy()#添加image
import json
import os
from PIL import Image
with open(os.path.join('./visium/CTR3_processed_data/spatial/scalefactors_json.json'), "r") as fh:
scales = json.load(fh)
img_hires = np.array(Image.open(os.path.join('./visium/CTR3_processed_data/spatial/tissue_hires_image.png')))
img_low = np.array(Image.open(os.path.join('./visium/CTR3_processed_data/spatial/tissue_lowres_image.png')))
#加入adata
lib_id = 'CTR3'
CTR3.uns["spatial"] = {
lib_id: {"images": {"hires": img_hires, "lowres": img_low}, "scalefactors": scales, "metadata": {}}
}CTR3AnnData object with n_obs × n_vars = 1801 × 18274
obs: 'orig.ident', 'nCount_Spatial', 'nFeature_Spatial', 'group', 'percent.mt', 'nCount_SCT', 'nFeature_SCT', 'nCount_RNA', 'nFeature_RNA', 'integrated_snn_res.0.6', 'seurat_clusters', 'in_tissue', 'array_row', 'array_col', 'pxl_row_in_fullres', 'pxl_col_in_fullres'
uns: 'seurat_clusters_colors', 'spatial'
obsm: 'pca', 'umap', 'spatial'sq.pl.spatial_scatter(CTR3)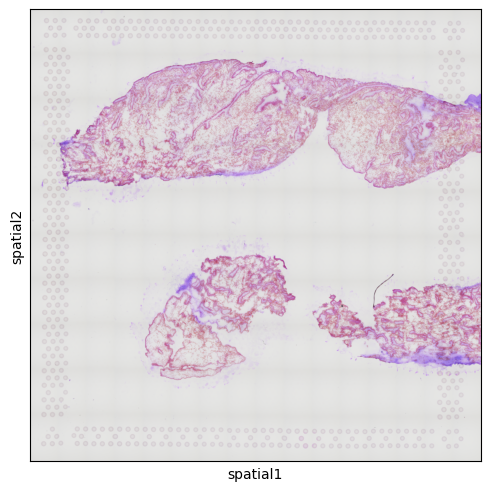
No description has been provided for this image
sc.pl.spatial(CTR3, img_key="lowres", color="seurat_clusters", size=1.5)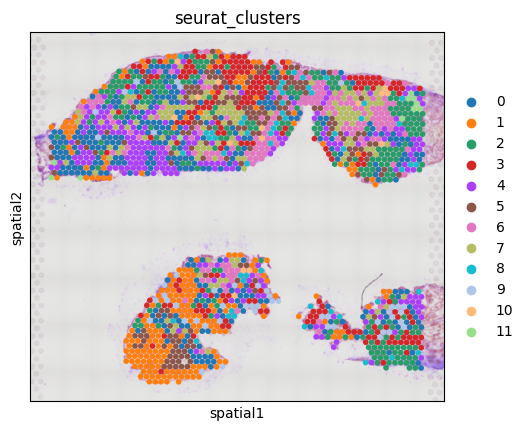
No description has been provided for this image
转化没有问题,但是很显然非常的麻烦!
所以,重点来了!
方法二:大家还是很期望一种直接转化的方式,一劳永逸,可以在转化对象的同时保留image,省去后续的麻烦。某些包是号称可以直接转化的,但是安装依赖太多,造成混乱,我尚未尝试,但是基本原理一样,都是类似于sceasy那样,调用python首先构建矩阵降维的转化,最后直接从seurat object获取image信息添加到h5ad。
这里参考seurat issue里面提供的一个函数(https://github.com/satijalab/seurat/issues/9617),这个函数大致方向是对的,但是最终也得不到需求的结果,而且有一些问题,直接无法使用。所以这里进行了修改重造,让其适用信更加广泛,实现空转visium seurat object转化为h5ad的“一步到位”实现。
看看我们函数的参数(函数需要提供image):对于scRNA数据,参数include_spatial选择F进行数据的转化,空转数据转化设置为T。
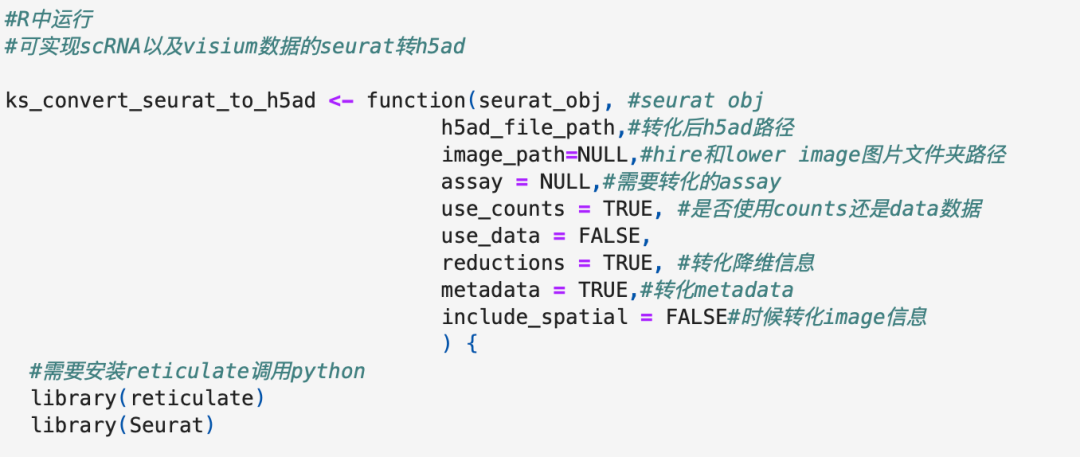

#转化演示
setwd("/home/ks_ts/data_analysis/Spatial_analysis/")
ks_convert_seurat_to_h5ad(seurat_obj = CTR3,h5ad_file_path = './',
image_path = "./visium/CTR3_processed_data/spatial/",
use_counts = FALSE,use_data = TRUE,
assay = 'SCT',include_spatial = T)#读入转化号的数据
CTR3 = sc.read_h5ad('./CTR3.h5ad')
CTR3
# AnnData object with n_obs × n_vars = 1801 × 18274
# obs: 'orig.ident', 'nCount_Spatial', 'nFeature_Spatial', 'group', 'percent.mt', 'nCount_SCT', 'nFeature_SCT', 'nCount_RNA', 'nFeature_RNA', 'integrated_snn_res.0.6', 'seurat_clusters'
# uns: 'spatial'
# obsm: 'pca', 'spatial', 'umap'可以看到,转化后的数据矩阵、降维、image信息都包括!
sq.pl.spatial_scatter(CTR3)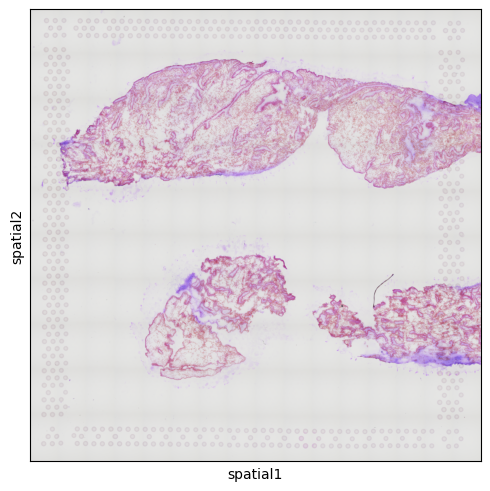
No description has been provided for this image
sq.pl.spatial_scatter(CTR3,color="seurat_clusters", size=1.5)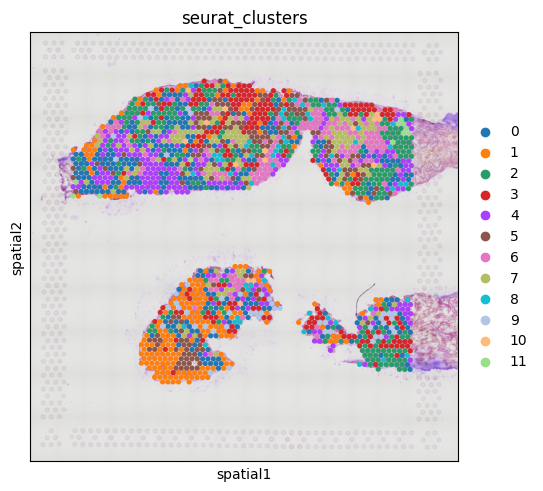
No description has been provided for this image
经过测试,可以实现。最后此函数算是一步到位的实现了。当然函数还有需要提高的地方,可以按照自己需求实现,让其功能更加强大!
觉得分享有用的点个赞再走呗!
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2026-03-15,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读
目录

